Attempting to get some sort of analysis of the Ostrea lurida GBS data from BGI, particularly since the last run at it using Stacks crashed (literally) and burned (not literally).
Katherine Silliman at UIC recommended using PyRAD.
I’ve taken the example Jupyter notebook from the PyRAD website and passed a subset of the 96 individuals through it.
In this instance, the subset of individuals were all analyzed as a single population. I have another Jupyter notebook running on a different computer that will separate the three populations that are present in this subset.
Overall, I don’t fully understand the results. However, this seems to be the quickest assessment of the data (from the *.snps file generated):
28 individuals, 36424 loci, 72251 snps
Additionally, I did run into an issue when I tried to visualize the data (using the *.vcf file generated) in IGV (see screen cap below). I’ve posted the issue to the pyrad GitHub repo in hopes of getting it resolved.
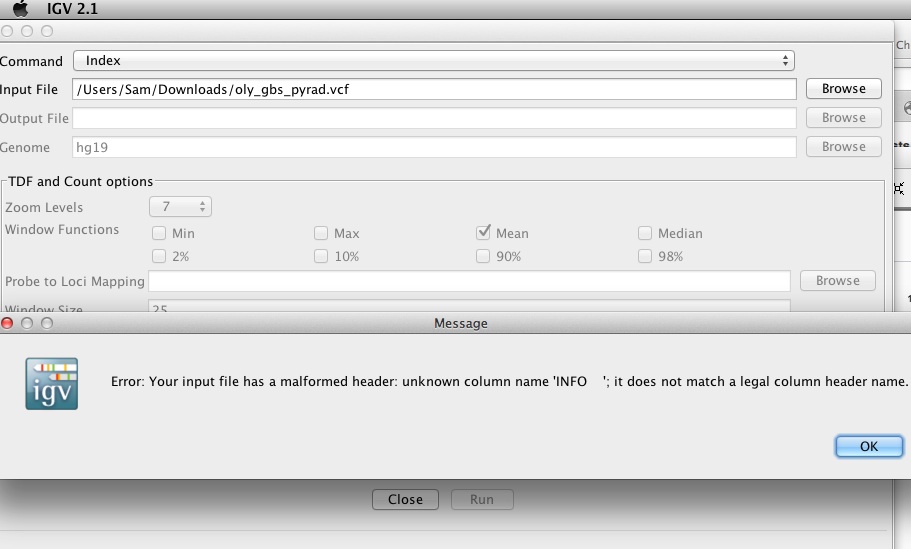 (http://eagle.fish.washington.edu/Arabidopsis/20160419_IGV_pyrad_vcf_index_error.jpg)
(http://eagle.fish.washington.edu/Arabidopsis/20160419_IGV_pyrad_vcf_index_error.jpg)
One last thing. This might be obvious to most, but I discovered that trying to do all this computation over the network (via a mounted server share) is significantly slower than performing these operations on th efiles when they’re stored locally. Somewhere in the notebook you’ll notice that I copy all of the working directory from the server (Owl) to the local machine (Hummingbird). Things proceeded very quickly after doing that. Didn’t realize this would have so much impact on speed!!
Jupyter Notebook: 20160418_pyrad_oly_PE-GBS.ipynb
NBviewer: 20160418_pyrad_oly_PE-GBS